Cotton Molecular Genetics
Our wild cotton germplasm collection contains representative accessions of all 17 Gossypium species indigenous to Australia. Two characteristics of the wild Australian cottons, the absence of insecticidal terpenoid aldehydes and resistance to the fungal wilt pathogen Fusarium oxysporum f. sp. vasinfectum, could contribute significantly to the improvement of elite cotton cultivars. Concurrently, this collection of wild Australian cotton species has been fundamental to ongoing biogeographic and evolutionary studies of the cotton genus (Gossypium).
Biogeography of Australian Gossypium Species

There are 17 Gossypium species indigenous to Australia. These species comprise three genetically and morphologically distinct groups recognized taxonomically as three sections within Gossypium subgenus Sturtia and cytogenetically as three genomes.
The C-genome (section Sturtia) contains Sturt's Desert Rose (G. sturtianum) and its western relative, G. robinsonii. Sturt's Desert Rose is the floral emblem of the Northern Territory but is found in seasonally dry creek beds in the temperate arid zones of all the mainland states. The more narrowly distributed G. robinsonii occurs only in the Pilbura region of Western Australia. Both species are large well-branched shrubs with mauve flowers with a deep maroon petal spot at the base of each petal.

The three G-genome species occur as low or spindly shrubs in the dry-monsoon and warm arid regions of Western Australia, Northern Territory, and Queensland. Gossypium australe is the most widespread and can occur in large populations along road verges. Gossypium bickii and G. nelsonii are mostly found in small populations in the Northern Territory and western Queensland. The G-genome species are sometimes known as Desert Roses because their flowers are very similar to those of Sturt's Desert Rose.

The 12 K-genome species occur in small populations in the monsoonal regions of the Kimberley in northern Western Australia. These herbaceous perennials are well adapted to the seasonal fires and droughts that characterize this region of Australia. During drought or fire the plant dies back to an underground rootstock and sends up new stems when conditions are suitable. K-genome species also possess a unique seed dispersal mechanism that is found no where else in the genus. The flowers are upright, but as the fruit develops, the stem curves toward the ground dropping the seed directly onto the ground when the fruit opens. The seeds have a fleshy appendage that attracts ants, which disperse the seed.
Scientific Staff: Brown, Brubaker, Craven
TOP
Gossypol Expression in Cotton

Cultivated cottons and their wild relatives produce a suite of chemicals known terpenoid aldehydes, commonly known as 'gossypol', that provide resistance to insect and disease pests. As a constitutive defence system, they are sequestered in lysigenous cavities that are found in virtually every part of the plant, including the seed. Microbial attack will elicit local terpenoid aldehyde synthesis. Unfortunately, the very toxicity that makes terpenoid aldehydes effective against pests also makes them toxic to non-ruminant animals.
Cottonseed-derived protein meals and oils are important by-products of cotton cultivation. However, these toxic terpenoid aldehydes must be removed before the seed oils can be used for human consumption or before the seed protein meals can be used as an animal feed for non-ruminants. The ideal cotton cultivar would not sequester terpenoid aldehydes in the seed, but would produce them as the seed germinates. The only wild cotton relatives that exhibit this pattern of terpenoid aldehyde expression are five species of Gossypium indigenous to Australia. The most well known of these species is the floral emblem of the Northern Territory, Sturt's Desert Rose (Gossypium sturtianum).

We are developing hybrid germplasm that incorporates chromosomes of the wild Australian species in a cultivated cotton background. These hybrid materials are useful in two ways. They provide a means of studying the genetic control of terpenoid expression in cotton, and may provide a means of transferring the relevant genes to a new generation of commercial cotton cultivars.
Scientific Staff: Brubaker
TOP
Resistance to Fusarium oxysporum f. sp. vasinfectum
The severity of Fusarium wilt (Fusarium oxysporum f. sp. vasinfectum) in cotton fields in Australia is rising with the number of affected fields and districts and the extent of infection increasing yearly. The Fusarium pathotypes responsible for this new challenge to the Australian cotton industry appear to be unique to Australia. Cotton cultivars resistant to Fusarium strains known from outside Australia are not resistant to these novel pathotypes (See: Fusarium spp. and Australian Native Cottons).
Because the native Australian cotton relatives may have evolved under exposure to these unique Fusarium strains, it is hoped that they will have useful resistance genes. In conjunction with the Queensland Department of Primary Industries and with funding from the Cotton Research and Development Corporation, we are testing geographically representative accessions of all the Australian Gossypium species for levels of and variation in resistance to Fusarium wilt. We hope that the Australian wild Gossypium species will contribute as sources of novel resistance genes, but also be model systems for clarifying the genetic control of Fusarium resistance. Knowing whether the resistance of the wild Australian cottons is conditioned by a single gene or is the cumulative effect of multiple genes is critical for developing sound breeding strategies, particularly those based on introgressing genes from wild crop relatives.
Scientific Staff: Becerra, Brubaker, Burdon, Wang
TOP
Cotton Genome Mapping
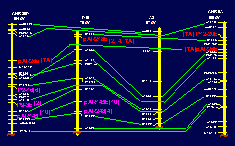
Genetic linkage maps are powerful tools in crop breeding and evolutionary studies. Currently most mapping efforts focus on cultivated crops, notwithstanding the utility of genetic linkage maps in studies of genome evolution and introgression. In cotton, genetic linkage maps have been useful characterising the conservation of gene order and synteny in the tetraploid Gossypium species relative to their diploid progenitors.
The Australian Gossypium species are distant relatives of the cultivated tetraploid cottons and their diploid progenitors. Comparable genetic linkage maps will provide a long-term perspective on genome evolution in the cotton genome.
Scientific Staff: Becerra, Brubaker
